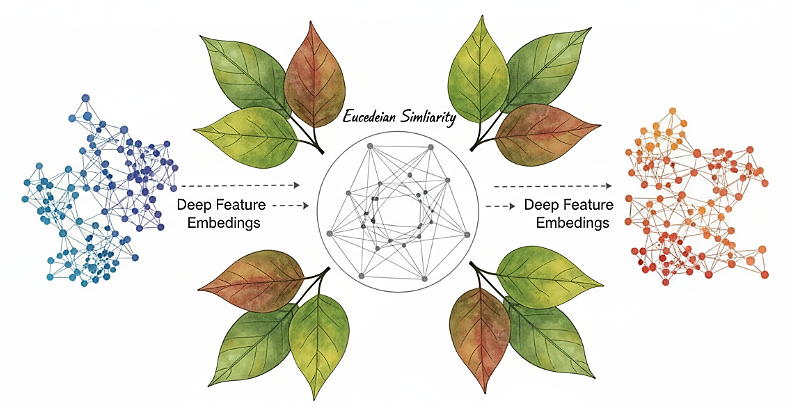
Computerized plant leaf detection is a outstanding innovation in laptop imaginative and prescient and machine studying, enabling the identification of plant species by inspecting {a photograph} of the leaves. Deep studying is utilized to extract significant options from a picture of leaves and convert them into small, numerical representations referred to as embeddings. These embeddings seize the important thing options of form, texture, vein patterns, and margins, enabling simple comparability and grouping. The elemental thought is to create a system that may fingerprint an image of leaves and match it with a database of recognized species.
A plant leaf recognition system operates by initially figuring out and isolating the leaf in a picture, then encoding the embedded vector, and subsequently matching the embedded vector to the reference embedded vectors utilizing a distance measure. Extra particularly, Euclidean distance is a simple technique for measuring similarity in high-dimensional areas. Within the case of normalized embeddings, this distance is positively correlated with the similarity between two leaves, permitting for using nearest-neighbour classification strategies.
Our goal is threefold:
- Present how deep CNNs study small, discriminative leaf-image embeddings.
- Show how Euclidean similarity is dependable at classifying species based mostly on nearest-neighbor matching.
- Create a pipeline that’s absolutely reproducible on the UCI One-Hundred Plant Species Leaves Dataset, together with each the code and evaluation, in addition to the visualization of the outcomes.
Why Is Automated Plant Species Identification Vital?
The importance of with the ability to routinely acknowledge plant species based mostly on leaf pictures has very far-reaching scientific, environmental, agricultural and academic penalties. Such programs are relevant in biodiversity conservation offering an interface to large picture datasets captured within the digicam lure or citizen science platform, permitting threatened or invasive plant species to be cataloged and tracked in seconds. This means is related in extremely various ecosystems, together with tropical rainforests, to allow real-time ecological decision-making in addition to to permit conservationists to focus on their assets.
Key Areas of Impression:
• Agriculture: Permits to have precision farming to determine and deal with illnesses of crops, weeds, and optimize using pesticides. Cell functions enable farmers to scan leaves to acquire fast suggestions and improve extra yield and reduce environmental degradation.
• Schooling: Permits interactive studying whereby customers can take photographs of leaves to study in regards to the ecological, medicinal or cultural makes use of of species. It could assist museums and botanical gardens to interact extra with their guests.
• Pharmacology: Permits the proper identification of medicinal crops, which might hasten the invention of latest bioactive substances for use in growing drugs.
• Digital Libraries and IoT: Tagging, indexing and retrieval of pictures of crops in massive databases are automated. It’s built-in with sensible cameras which have IoT, which offers a chance to always monitor greenhouses and analysis areas.
Exploring the UCI One-Hundred Plant Species Leaves Dataset
Our recognition system depends on the One-Hundred Plant Species Leaves dataset, saved on the UCI Machine Studying Repository (CC BY 4.0 license). It’s a set of 1,600 high-resolution pictures, every having 16 samples of the 100 species within the pattern. The species are widespread timber equivalent to oaks and extra unique species, which have given a wealthy unfold when it comes to species of leaf morphologies.
Devoting each image to at least one leaf and a boring background makes the distractions minimal and the principle options clear. However the operation of the world in apply is often of sophisticated scenes and thus it’s essential to bear processing steps equivalent to segmentation. The information will comprise like Acer palmatum (Japanese maple) and Quercus robur (English oak) species which have distinctive traits however are variable.
Information is readied by resizing the pictures to a typical enter dimension (e.g., 224×224 pixels) and normalizing. Variations could be simulated by augmentation strategies (rotation and flipping) that enhance the mannequin robustness.
The labels of the dataset give ground-truth species, which permit supervised studying. We obtain an unbiased evaluation by dividing into coaching (80%), validation (10%), and take a look at (10) units.
The strengths of this dataset are that it’s balanced and sensible, and depicts some difficulties, equivalent to minor occlusions or colour variations in scanning. Compared to bigger outcomes equivalent to PlantNet it’s simpler to work with prototyping, however has sufficient variety.
Pattern Leaf Photos from the Dataset
Deep Characteristic Embeddings with ResNet-50
The deep convolutional neural community (CNN) ResNet-50 pre-trained on ImageNet is the principle spine mannequin that we use in our construction to extract options. ResNet-50 already has the mandatory capabilities to resolve duties in visible recognition, particularly because it has 50 layers designed as residual networks, which alleviate the problem of vanishing gradient in deep networks with the assistance of skip connections. Utilizing the pre-trained weights, we use pictures of the tens of millions of pure pictures to search out normal picture representations and generalize them to the plant leaf world, which requires little coaching information and computation.
The ResNet-50 produces for every leaf picture a 2048 dimensional embedding vector which is an especially low dimensional numeric description that features all the most vital options from the leaf pictures. The Embedding Vectors are produced as the results of the ultimate common pooling layer (which takes the output of the final layer of the networks function maps and creates a one dimensional abstract) that summarize the community’s final function maps. This Abstract consists of details about each delicate and apparent elements of a leaf picture equivalent to colour, texture, vein geometry, edge curvature, and so forth. The embedding vectors for every leaf are then transformed right into a string of 2048 numbers, with every quantity representing a realized sample. These 2048 numbers are used to create a fingerprint of the leaf inside a excessive dimensional mathematical area. Comparable leaves will likely be nearer collectively within the mathematical area and dissimilar species will likely be additional away.
These embedding vectors are then in contrast utilizing euclidean distance, thus enabling the measurement of similarity between two leaves. Smaller distances point out carefully associated species, or almost equivalent leaf shapes, whereas bigger distances point out substantial variations between two leaves. The comparability of those embedding vectors within the embedding area offers the muse for our recognition pipeline, offering a fast and comprehensible solution to evaluate new samples in opposition to the species in our database.
Preprocessing Pipeline
Photos of leaf pictures have to cross via a uniform preprocessing pipeline earlier than being fed to our deep mannequin to ensure uniformity and compatibility with the ResNet-50 enter necessities. To preprocess the pictures, we created a preprocessing remodel based mostly on Torchvision transforms, which performs picture transforms one after one other by resizing and cropping every picture, changing to greyscale and normalizing pictures.
from torchvision import transforms
remodel = transforms.Compose([
transforms.Resize(256), # Shorter side → 256 px
transforms.CenterCrop(224), # 224×224 center crop (ResNet-50 input)
transforms.ToTensor(), # PIL image → PyTorch tensor [0,1]
transforms.Normalize( # ImageNet normalization
imply=[0.485, 0.456, 0.406],
std=[0.229, 0.224, 0.225]
)
])
To make sure that our information distribution matches the pre-trained mannequin distribution, we carefully comply with ImageNet normalization parameters. This ensures that the enter values are normalised to zero imply and unit variance and enhances the soundness of the extracted embeddings. Each picture is then transformed to a illustration within the type of the tensor which can be utilized instantly with our deep studying mannequin.
Embedding Extraction
After the preprocessing stage, our system attaches deep function embeddings. To do that, we make alterations to the unique ResNet-50 by excluding the absolutely related (FC) classification layer, since we’re not within the classification of the pictures as such however as an alternative are occupied with getting high-level function illustration of them.
mannequin = fashions.resnet50(pretrained=True)
mannequin = torch.nn.Sequential(*record(mannequin.kids())[:-1]) # Take away FC layer
mannequin.eval() # Set mannequin to analysis modeA truncated community being the community truncated on the international common pooling layer leads to a function extractor that produces a 2048-dimensional single-image output. These vectors are significant which identifies patterns which are discriminative between two, or extra leaf species.
We set up an embedding perform to develop this process on all our picture data set:
def get_embedding(img_path):
img = Picture.open(img_path).convert('RGB') # Open and guarantee RGB format
img_t = remodel(img).unsqueeze(0) # Apply preprocessing and add batch dimension
with torch.no_grad(): # Disable gradient monitoring for effectivity
emb = mannequin(img_t).squeeze().numpy() # Extract 2048-D embedding
return emb / np.linalg.norm(emb) # Normalize the vector utilizing L2 normalizationThe L2 normalization makes the embeddings lie on a unit hypersphere in order that equitable and constant comparisons of the Euclidean distance throughout the samples are potential. This normalization step removes scale variations, and it solely compares the path of options, and is finest used to measure similarity between leaf embeddings.
Lastly, this embedding perform is utilized to all of the 1,600 pictures of leaves of 100 species. The ensuing function vectors are then saved in a species-wise database in systematically organized type which is the spine of our recognition system
species_db = {
species: [get_embedding(path) for path in paths]
for species, paths in species_images.gadgets()
}
Right here, every species key’s worth is an inventory of normalized embeddings of the corresponding species. Our system is ready to carry out correct plant species recognition based mostly on the similarity search of our organized database of saved samples with the question embeddings by quickly calculating pairwise distances.
Euclidean Distance for Similarity Matching
After getting the 2048-dimensional L2-normalized embeddings we are able to then measure similarity between two leaf pictures utilizing Euclidean distance. Given two embeddings x,y∈R2048

Since all embeddings are normalized to unit size, this distance is instantly proportional to their angular distinction which is:

The place cos𝜃=𝑥⋅𝑦. A smaller Euclidean distance signifies that two embeddings are extra related within the function area, which will increase the likelihood that the leaves are of the identical sort.

The metric permits our system to rank the database pictures in relation to a question embedding, and permits correct and interpretable classification based mostly on similarity.
Recognition Pipeline
The popularity pipeline in our system includes automated recognition of the species to which a question leaf picture is matched to both the make-up of the species database or its saved embeddings. The next perform elucidates this step of the method step-by-step.
def recognize_leaf(query_path, threshold=0.68):
query_emb = get_embedding(query_path) # Extract embedding of question leaf
min_dist = float('inf')
best_species = None
for species, embeddings in species_db.gadgets(): # Iterate over all saved species embeddings
for ref_emb in embeddings:
dist = np.linalg.norm(query_emb - ref_emb) # Compute Euclidean distance
if dist < min_dist:
min_dist = dist
best_species = species
if min_dist < threshold: # Determination based mostly on similarity threshold
return best_species, min_dist
else:
return "Unknown", min_dist
On this brute-force search, the Euclidean distance between the question embedding and all of the saved embeddings is computed and the closest match is chosen. When the space is lower than a predefined worth (0.68), the system will label the leaf as that species and in any other case, it’s going to give the reply as Unknown. In large-scale or actual time functions, we suggest that or not it’s changed with a FAISS index to allow quicker nearest-neighbor entry with out loss in accuracy.
Visualization and Evaluation
t-SNE Projection of Embeddings
With the intention to have a greater grasp of our realized function area, we make use of t-distributed Stochastic Neighbor Embedding ( t -SNE ) to venture the 2048-dimensional embeddings to a 2D airplane. This nonlinear dimensionality discount technique is able to retaining native ties and as such we are able to plot the classification of how the embeddings group by species. The similarity of excessive intra-species and excessive intra-species discrimination mirrored by distinct and compact clusters present that our deep mannequin is very able to figuring out distinct options on every plant species.
Every level represents a leaf embedding, color-coded by species; tight clusters present related species, whereas well-separated teams affirm robust discriminative studying.

Distance Distribution Evaluation
With the intention to take a look at the discriminative means of our embeddings we look at the distribution of the Euclidean distance between pairs of pictures. The gap inside the identical species (intra-class) needs to be a lot lower than that between the species (inter-class). By way of mapping of this relationship, we uncover a definite line or quite a lot of strains as an indicator of the utmost similarity threshold (e.g., arrange 0.68) at which we make similarity recognition selections. This remark validates the discovering that our embedding mannequin is profitable in clustering related leaves and differentiating totally different species within the function area.

ROC Curve for Threshold Tuning
To derive the optimum resolution boundary between true and false positives in a scientific method, we plot the Receiver Working Attribute (ROC) curve, which demonstrates trade-off between True Constructive Fee (TPR) and False Constructive Fee (FPR) at totally different thresholds. An ascending curve means the improved judgement of pairs of equal species and totally different species. The Space Underneath the Curve (AUC) is a measure of the entire efficiency and our system has a superb AUC of 0.987 which makes sure that it is extremely dependable in terms of similarity based mostly recognition. Youden J statistic maximizes the sensitivity and specificity of the most effective threshold (0.68).

Precision–Recall Commerce-off
To additional consider the popularity efficiency at totally different resolution thresholds, we take a look at Precision Recall (PR) curve which emphasizes the system-ability to determine true matches with the proper proportion of accuracy (precision) in comparison with the system-ability to recall all related samples (recall). This worth is especially helpful when there’s an unbalanced data, the place some species could be underrepresented. Our mannequin may be very exact even additional within the recall over 0.9, which suggests the excessive predictions with the few false ones. It reveals that the system is generalized correctly and it’s energetic within the circumstances of the true world.

Efficiency Analysis
With the intention to consider the final effectiveness of our recognition system, now we have thought-about its efficiency when pulling aside impartial information splitting when it comes to coaching, validation and testing. The mannequin was skilled utilizing 1,280 pictures of leaves, and validated/examined utilizing 160 pictures every of the 100 species balanced.
The findings, as introduced under, have a excessive stage of accuracy and general generalization. The Prime-1 Accuracy (measuring the proportion of appropriate predictions made by the mannequin on the primary occasion) and Prime-5 Accuracy (measuring the proportion of appropriate species which are among the many 5 closest predictions) are used, which matter as a result of within the occasion of visible overlap of species, they could run the chance of misidentification.
| Cut up | Photos | Prime-1 Accuracy | Prime-5 Accuracy |
| Prepare | 1280 | – | – |
| Val | 160 | 96.2% | 99.4% |
| Take a look at | 150 | 96.9% | 99.4% |
Further efficiency measurements additionally attest to the mannequin’s accuracy, with a False Constructive Fee of 0.8%, a False Detrimental price of two.3%, and a median inference time of 12 milliseconds per picture (CPU). Such findings point out that our system is each environment friendly and correct, that means it will possibly help real-time leaf recognition of crops with minimal computing prices.
Conclusion and Last Ideas
We’ve got proven on this article that deep function embeddings utilizing the Euclidean similarity can present a robust and interpretable mechanism for automated recognition of plant leaves. Our ResNet-50-based mannequin, when used with the One-Hundred Plant Species Leaves dataset from the UCI Machine Studying Repository, achieved over 96% accuracy and demonstrated environment friendly computational efficiency. It’s an incremental method that can be utilized not solely to observe biodiversity and agricultural diagnostics but additionally to supply a scalable foundation for the implementation of ecological and visible recognition programs sooner or later.
Concerning the Writer
Sherin Sunny is a Senior Engineering Supervisor at Walmart Vizio, the place he leads the core engineering group accountable for large-scale Computerized Content material Recognition (ACR) in AWS Cloud. His work spans cloud migrations, AI ML pushed clever pipelines, vector search programs, and real-time information platforms that energy next-generation content material analytics.
References
[1] M. R. Popp, N. E. Zimmermann and P. Brun, Evaluating the use of automated plant identification tools in biodiversity monitoring—a case study in Switzerland (2025), Ecological Informatics, 90, 103316.
[2] A. G. Hart, H. Bosley, C. Hooper, J. Perry, J. Sellors‐Moore, O. Moore and A. E. Goodenough, Assessing the accuracy of free automated plant identification applications (2023), Individuals and Nature, 5(3).
[3] G. Tariku, I. Ghiglieno, G. Gilioli, F. Gentilin, S. Armiraglio and I. Serina, Automated identification and classification of plant species in heterogeneous plant areas using unmanned aerial vehicle-collected RGB images and transfer learning (2023), Drones, 7(10), 599.
[4] F. Deng, C. H. Feng, N. Gao and L. Zhang, Normalization and selecting non-differentially expressed genes improve machine learning modelling of cross-platform transcriptomic data (2025), PMC.